|
Size: 267
Comment:
|
Size: 5137
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| ## page was renamed from User's Guide ## page was renamed from User Guide |
|
| Line 7: | Line 9: |
| Line 9: | Line 10: |
| Line 16: | Line 15: |
| == How to run BDF == To run BDF, you can write a shell script named "run.sh" with following content, {{{ #!/bin/bash # Set BDF home directory export BDFHOME=~/work/0.5.dev # run BDF driver with input file $1 $BDFHOME/sbin/bdfdrv.py -r $1 }}} For example, you can copy the file named "$BDFHOME/Tests/input/test002.inp" to a work directory. Then, you write done the shell script and store it in you work directory. To evoke BDF calculation, you just use command {{{ $./run.sh test001.inp }}} The output will be printed on standard output. Thus, it is better to redirect output to a file. {{{ $./run.sh test001.inp > test001.out }}} == Some tips to run BDF == {{{ 1. There are a lot of testing inputs saved in directory of $BDFHOME/Tests/input. 2. BDF driver assume input file has the name *.inp. Thus, you can run BDF with command $./run.sh test001 3. If BDF is compiled with OpenMP supporting, you can can set OpenMP environmental variables in running script. For example, export OMP_NUM_THREADS=4 export OMP_STACKSIZE=1024M 4. You can use shell command in BDF input files. For example, you can backup HF canonical orbitals after SCF calculation. $SCF $END %cp $BDF_WORKDIR/$BDFTASK.scforb $BDF_WORKDIR/myscforb.bak }}} == BDF Flowchat == {{attachment:bdf_module_chart.jpg||width=640,align="middle"}} == Input style == == Environmental variables used in BDF == There are some important environmental variables used in BDF. {{{ 1.BDF_WORKDIR - Work directory used in running BDF program. 2.BDF_TMPDIR - Scratch directory used in running BDF program. This directory can be removed after the calculation. 3.BDFTASK - Task name of a BDF work. }}} == BDF modules == [[compass]] - Molecule geometry and basis set preprocess. [[drt]] - Generate DRTs in GUGA. [[grad]] - Gradient. [[mcscf]] - Multi-configuration self-consistent-field program [[mp2]] - MP2 program [[mrci]] - Multi-reference configuration interaction program. [[localmo]] - Localization of molecule orbital. [[scf]] - Self-consistent-field program. [[tddft]] - Time dependent density functional program. [[vgmfci]] - electron-nucleus mean field configuration interaction program. [[xuanyuan]] - 1e and 2e integrals program. [[genfrag]] - Generate or optimize fragments and fragments pairs in Local orbital based Frag-MP2/CCSD. [[expandmo]] - Expand molecular orbital from small basis set to large basis set. [[resp]] - Module for response properties based on HF and DFT == QM/MM calculation == [[External charge]] -- Input point charges. A file named $BDFTASK.extcharg should be prepared. Here is an example. [[H2O.inp]] {{{ $COMPASS Title water molecule in backgroud of exteral charges Basis 6-31g Geometry O 0.000000 0.000000 0.106830 H 0.000000 0.785178 -0.427319 H 0.000000 -0.785178 -0.427319 End Geometry Extcharge # required if external charge exists. point # required if external charge exists. "point" specifies the type of external charge is point charge. Check Skeleton $END $XUANYUAN direct schwarz $END $SCF RHF $END }}} [[H2O.extcharge]] {{{ External charge, Point charge # title line 6 # number of point charges. Next six lines are label, charge, x,y,z coordinates. Unit: angstrom(default) C1 -0.732879 0.000000 5.000000 0.114039 C2 0.366440 0.000000 5.780843 -0.456155 C3 0.366440 0.000000 4.219157 -0.456155 C4 -0.732879 0.000000 10.00000 0.114039 C5 0.366440 0.000000 10.78084 -0.456155 C6 0.366440 0.000000 9.219157 -0.456155 }}} Another example {{{ External charge, Point charge # title line 6 Bohr # number of point charges, Unit: Bohr C1 -0.732879 0.000000 5.000000 0.114039 C2 0.366440 0.000000 5.780843 -0.456155 C3 0.366440 0.000000 4.219157 -0.456155 C4 -0.732879 0.000000 10.00000 0.114039 C5 0.366440 0.000000 10.78084 -0.456155 C6 0.366440 0.000000 9.219157 -0.456155 }}} == PCM solver == To use PCM in BDF, you need add PCMSOLVER in SCF input. Then a pcminput section is required if you would like change default parameters in PCM solver. {{{ $SCF ... PCMSOLVER # require PCM module $END &PCMINPUT # PCM input Solvent Chloroform # Set solvent as CHCl3 &END For detail: please check [[PCMinput]] parameters. }}} |
|
| Line 18: | Line 169: |
| == BDF modules == | == Useful tools == [[bdffrag]] |
BDF User's guide
Contents
Insert introduction of BDF module at here.
How to run BDF
To run BDF, you can write a shell script named "run.sh" with following content,
#!/bin/bash # Set BDF home directory export BDFHOME=~/work/0.5.dev # run BDF driver with input file $1 $BDFHOME/sbin/bdfdrv.py -r $1
For example, you can copy the file named "$BDFHOME/Tests/input/test002.inp" to a work directory. Then, you write done the shell script and store it in you work directory. To evoke BDF calculation, you just use command
$./run.sh test001.inp
The output will be printed on standard output. Thus, it is better to redirect output to a file.
$./run.sh test001.inp > test001.out
Some tips to run BDF
1. There are a lot of testing inputs saved in directory of $BDFHOME/Tests/input.
2. BDF driver assume input file has the name *.inp. Thus, you can run BDF with command
$./run.sh test001
3. If BDF is compiled with OpenMP supporting, you can can set OpenMP environmental variables in running script. For example,
export OMP_NUM_THREADS=4
export OMP_STACKSIZE=1024M
4. You can use shell command in BDF input files. For example, you can backup HF canonical orbitals after SCF calculation.
$SCF
$END
%cp $BDF_WORKDIR/$BDFTASK.scforb $BDF_WORKDIR/myscforb.bak
BDF Flowchat
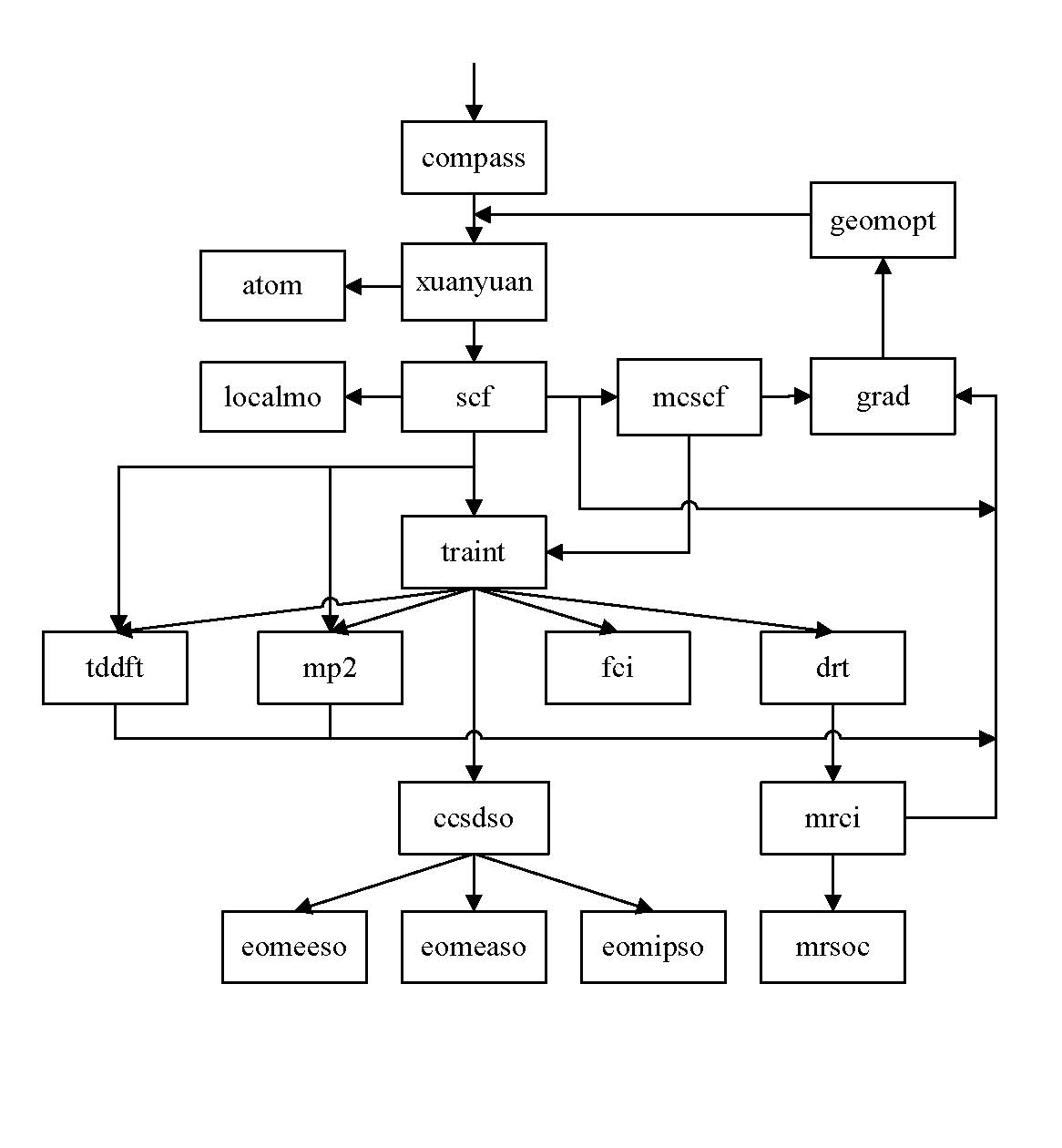
Input style
Environmental variables used in BDF
There are some important environmental variables used in BDF.
1.BDF_WORKDIR - Work directory used in running BDF program. 2.BDF_TMPDIR - Scratch directory used in running BDF program. This directory can be removed after the calculation. 3.BDFTASK - Task name of a BDF work.
BDF modules
compass - Molecule geometry and basis set preprocess.
drt - Generate DRTs in GUGA.
grad - Gradient.
mcscf - Multi-configuration self-consistent-field program
mp2 - MP2 program
mrci - Multi-reference configuration interaction program.
localmo - Localization of molecule orbital.
scf - Self-consistent-field program.
tddft - Time dependent density functional program.
vgmfci - electron-nucleus mean field configuration interaction program.
xuanyuan - 1e and 2e integrals program.
genfrag - Generate or optimize fragments and fragments pairs in Local orbital based Frag-MP2/CCSD.
expandmo - Expand molecular orbital from small basis set to large basis set.
resp - Module for response properties based on HF and DFT
QM/MM calculation
External charge -- Input point charges. A file named $BDFTASK.extcharg should be prepared. Here is an example.
$COMPASS Title water molecule in backgroud of exteral charges Basis 6-31g Geometry O 0.000000 0.000000 0.106830 H 0.000000 0.785178 -0.427319 H 0.000000 -0.785178 -0.427319 End Geometry Extcharge # required if external charge exists. point # required if external charge exists. "point" specifies the type of external charge is point charge. Check Skeleton $END $XUANYUAN direct schwarz $END $SCF RHF $END
External charge, Point charge # title line 6 # number of point charges. Next six lines are label, charge, x,y,z coordinates. Unit: angstrom(default) C1 -0.732879 0.000000 5.000000 0.114039 C2 0.366440 0.000000 5.780843 -0.456155 C3 0.366440 0.000000 4.219157 -0.456155 C4 -0.732879 0.000000 10.00000 0.114039 C5 0.366440 0.000000 10.78084 -0.456155 C6 0.366440 0.000000 9.219157 -0.456155
Another example
External charge, Point charge # title line 6 Bohr # number of point charges, Unit: Bohr C1 -0.732879 0.000000 5.000000 0.114039 C2 0.366440 0.000000 5.780843 -0.456155 C3 0.366440 0.000000 4.219157 -0.456155 C4 -0.732879 0.000000 10.00000 0.114039 C5 0.366440 0.000000 10.78084 -0.456155 C6 0.366440 0.000000 9.219157 -0.456155
PCM solver
To use PCM in BDF, you need add PCMSOLVER in SCF input. Then a pcminput section is required if you would like change default parameters in PCM solver.
$SCF ... PCMSOLVER # require PCM module $END &PCMINPUT # PCM input Solvent Chloroform # Set solvent as CHCl3 &END For detail: please check [[PCMinput]] parameters.