|
Size: 5582
Comment:
|
Size: 5645
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 41: | Line 41: |
| 3. If BDF is compiled with OpenMP supporting, you can can set OpenMP environmental variables in running script. For example, | 3. If BDF is compiled with OpenMP supporting, you can can set OpenMP environment variables in running script. For example, |
| Line 48: | Line 48: |
| 5. A sample run.sh file is available in bdf-pkg/sbin/run.sh. | |
| Line 70: | Line 71: |
[[ bdfopt ]] - Molecular geometry optimizer. |
|
| Line 73: | Line 77: |
[[ Elecoup ]] - Electron transfer integral, electric excited states coupling etc. [[expandmo]] - Expand molecular orbital from the small basis set to large basis set. [[genfrag]] - Generate or optimize fragments and fragments pairs in Local orbital based Frag-MP2/CCSD. |
|
| Line 92: | Line 102: |
| [[genfrag]] - Generate or optimize fragments and fragments pairs in Local orbital based Frag-MP2/CCSD. [[expandmo]] - Expand molecular orbital from the small basis set to large basis set. |
|
| Line 97: | Line 103: |
[[xianci]] - Module for MRCISD and MRPT2 calculations. Interfaced with the Xian-CI package [[ bdfopt ]] - Molecular geometry optimizer. |
|
| Line 104: | Line 106: |
| [[ Elecoup ]] - Electron transfer integral, electric excited states coupling etc. | [[xianci]] - Module for MRCISD and MRPT2 calculations. Interfaced with the Xian-CI package |
BDF User's guide
Contents
How to run BDF
To run BDF, you can write a shell script named "run.sh" with following content,
#!/bin/bash # Set BDF home directory export BDFHOME=~/work/0.5.dev # run BDF driver with input file $1 $BDFHOME/sbin/bdfdrv.py -r $1
For example, you can copy the file named "$BDFHOME/Tests/input/test002.inp" to a work directory. Then, you write down the shell script and store it in you work directory. To evoke BDF calculation, you just use command
$./run.sh test001.inp
The output will be printed on standard output. Thus, it is better to redirect output to a file.
$./run.sh test001.inp > test001.out
Some tips to run BDF
1. There are a lot of testing inputs saved in directory of $BDFHOME/Tests/input.
2. BDF driver assume input file has the name *.inp. Thus, you can run BDF with command
$./run.sh test001
3. If BDF is compiled with OpenMP supporting, you can can set OpenMP environment variables in running script. For example,
export OMP_NUM_THREADS=4
export OMP_STACKSIZE=1024M
4. You can use shell command in BDF input files. For example, you can backup HF canonical orbitals after SCF calculation.
$SCF
$END
%cp $BDF_WORKDIR/$BDFTASK.scforb $BDF_WORKDIR/myscforb.bak
5. A sample run.sh file is available in bdf-pkg/sbin/run.sh.
BDF easy input
BDF easy input is in developing ... bdfeasyinput
BDF Flowchart
Note this is the execution order of the modules, which is usually, but not always, the same as the order of input blocks in the input file.
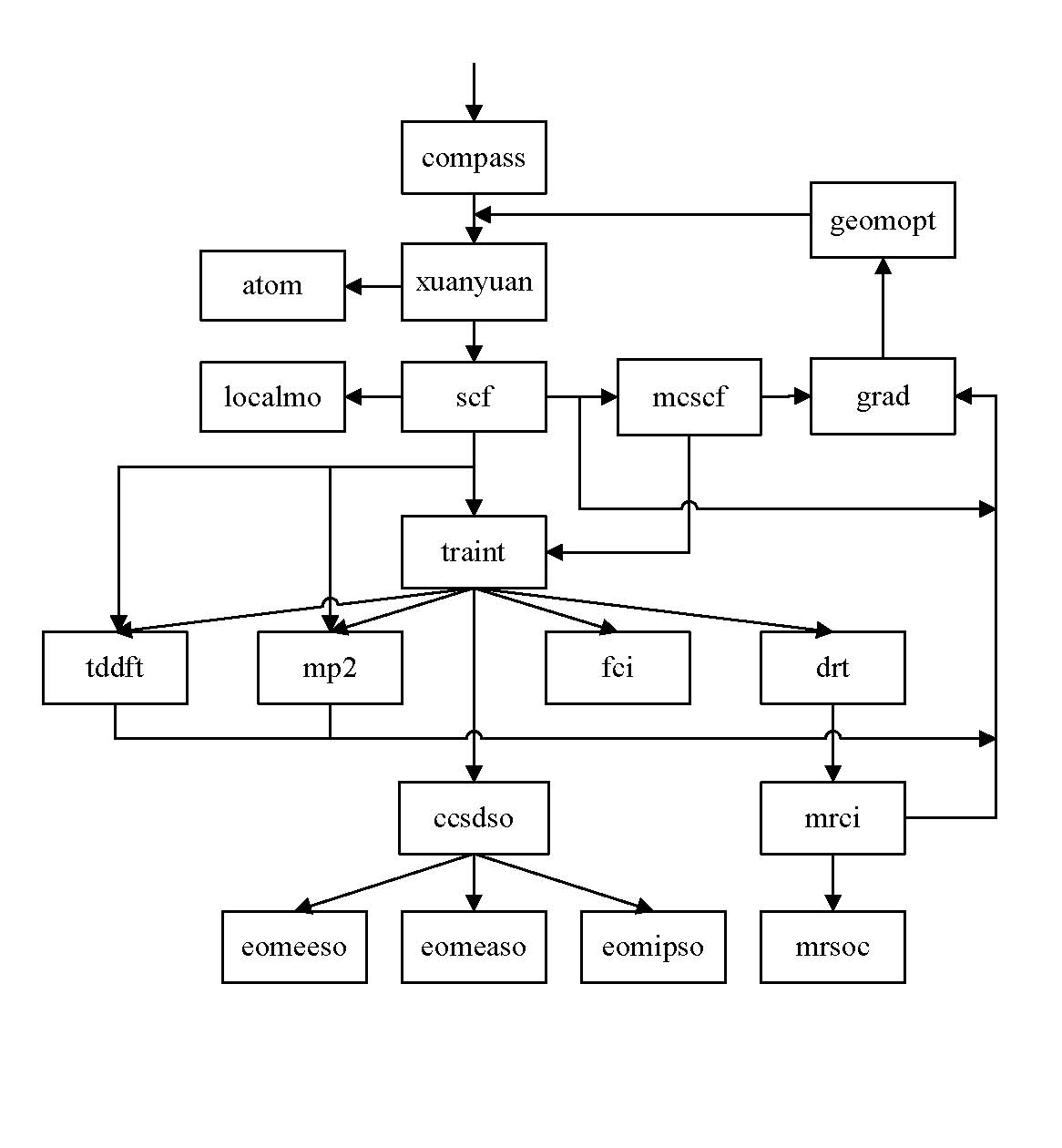
Input style
Environmental variables used in BDF
There are some important environmental variables used in BDF.
1.BDF_WORKDIR - Work directory used in running BDF program. 2.BDF_TMPDIR - Scratch directory used in running BDF program. This directory can be removed after the calculation. 3.BDFTASK - Task name of a BDF work.
BDF modules
bdfopt - Molecular geometry optimizer.
compass - Molecule geometry and basis set preprocess.
drt - Generate DRTs in GUGA.
Elecoup - Electron transfer integral, electric excited states coupling etc.
expandmo - Expand molecular orbital from the small basis set to large basis set.
genfrag - Generate or optimize fragments and fragments pairs in Local orbital based Frag-MP2/CCSD.
grad - Gradient.
mcscf - Multi-configuration self-consistent-field program
mp2 - MP2 program
mrci - Multi-reference configuration interaction program.
localmo - Localization of molecule orbital.
scf - Self-consistent-field program.
tddft - Time dependent density functional program.
vgmfci - electron-nucleus mean field configuration interaction program.
xuanyuan - 1e and 2e integrals program.
resp - Module for response properties based on HF and DFT
Traint - AO to MO integral transformation.
xianci - Module for MRCISD and MRPT2 calculations. Interfaced with the Xian-CI package
QM/MM calculation
External charge -- Input point charges. A file named $BDFTASK.extcharge should be prepared. Here is an example.
$COMPASS Title water molecule in backgroud of exteral charges Basis 6-31g Geometry O 0.000000 0.000000 0.106830 H 0.000000 0.785178 -0.427319 H 0.000000 -0.785178 -0.427319 End Geometry Extcharge # required if external charge exists. point # required if external charge exists. "point" specifies the type of external charge is point charge. Check Skeleton $END $XUANYUAN direct schwarz $END $SCF RHF $END
External charge, Point charge # title line 6 # number of point charges. Next six lines are label, charge, x,y,z coordinates. Unit: angstrom(default) C1 -0.732879 0.000000 5.000000 0.114039 C2 0.366440 0.000000 5.780843 -0.456155 C3 0.366440 0.000000 4.219157 -0.456155 C4 -0.732879 0.000000 10.00000 0.114039 C5 0.366440 0.000000 10.78084 -0.456155 C6 0.366440 0.000000 9.219157 -0.456155
Another example
External charge, Point charge # title line 6 Bohr # number of point charges, Unit: Bohr C1 -0.732879 0.000000 5.000000 0.114039 C2 0.366440 0.000000 5.780843 -0.456155 C3 0.366440 0.000000 4.219157 -0.456155 C4 -0.732879 0.000000 10.00000 0.114039 C5 0.366440 0.000000 10.78084 -0.456155 C6 0.366440 0.000000 9.219157 -0.456155
PCM solver
To use PCM in BDF, you need add PCMSOLVER in SCF input. Then a pcminput section is required if you would like to change the default parameters in PCM solver. For detail: please check PCMinput parameters.
$SCF ... PCMSOLVER # require PCM module $END &PCMINPUT # PCM input Solvent Chloroform # Set solvent as CHCl3 &END